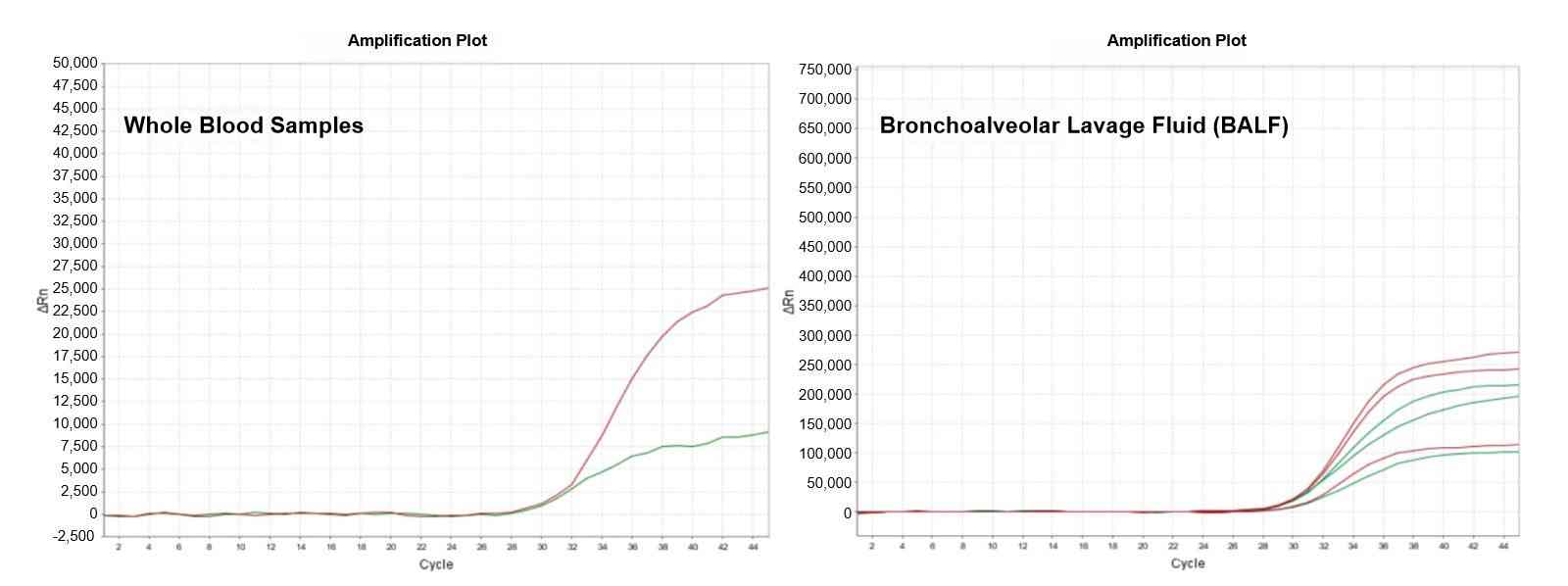
ENZD-P06 was evaluated for methylation-specific qPCR using bisulfite-converted DNA templates under standardized reaction conditions. Performance was compared with commercially available methylation qPCR master mixes.
The results demonstrated improved amplification linearity and higher fluorescence plateau signals, indicating enhanced reaction efficiency and more stable signal output in methylation detection workflows. These results are derived from internal validation studies evaluating amplification performance in bisulfite-converted DNA qPCR systems.
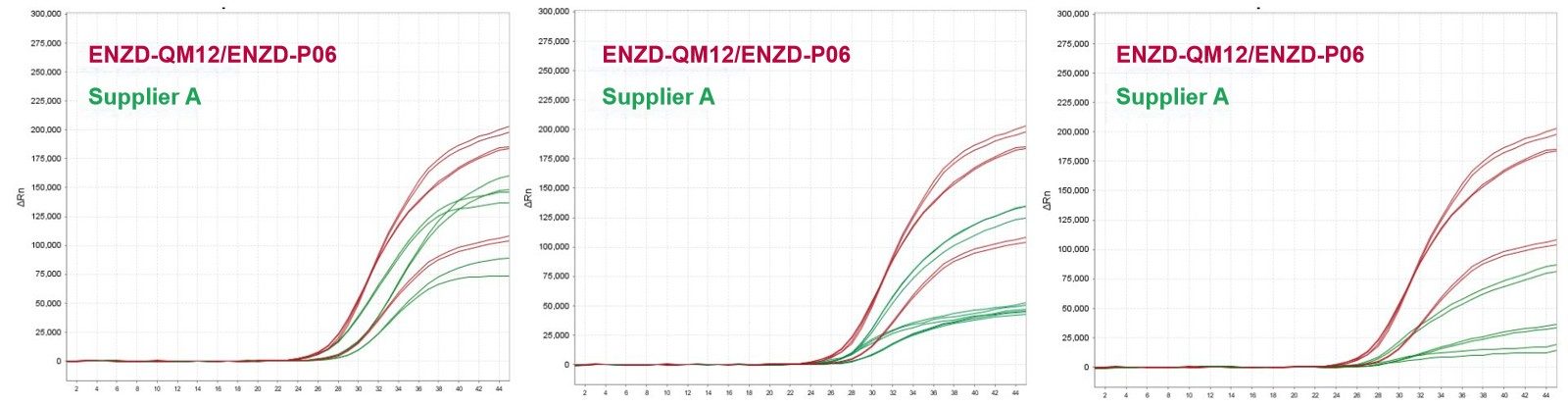
CpG islands within bisulfite-converted DNA often exhibit wide GC content variation, ranging from low to high GC regions, which can introduce amplification bias during methylation analysis.
ENZD-P06 demonstrated robust performance across templates with GC content ranging from 20% to 75%, ensuring consistent amplification efficiency across diverse genomic regions. Compared with Supplier D, ENZD-P06 showed reduced GC-dependent bias and improved amplification uniformity. These findings are based on internal validation studies assessing GC-content dependent amplification performance across 20%–75% GC templates.
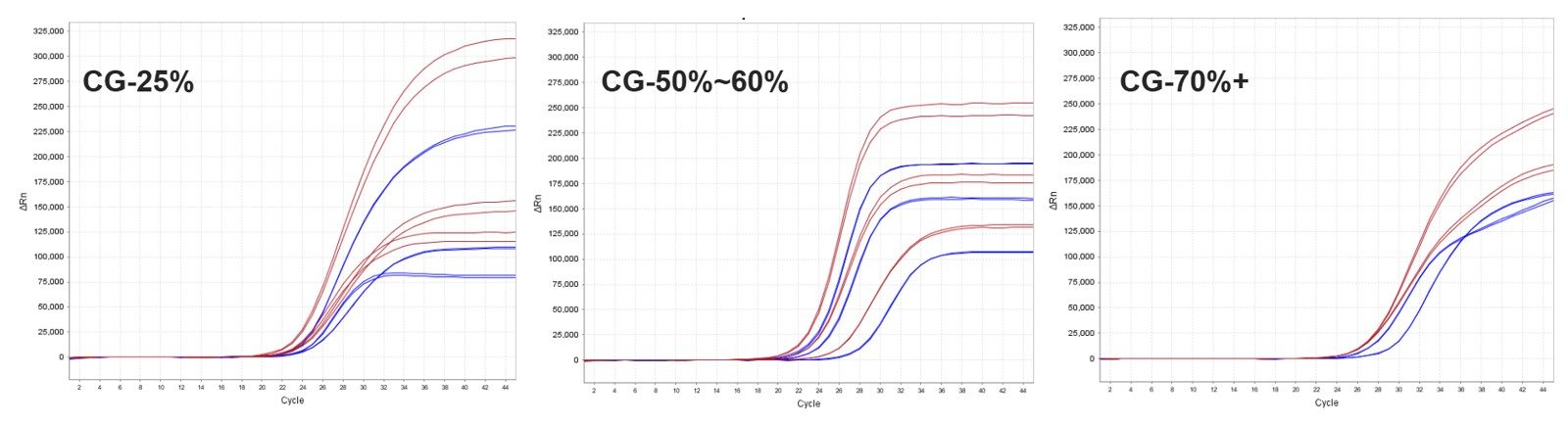
The performance of ENZD-P06 was evaluated using bisulfite-converted DNA derived from multiple clinically relevant sample types, including fecal extracts, exfoliated epithelial cells, blood-derived samples, and bronchoalveolar lavage fluid.
ENZD-P06 demonstrated robust amplification performance across all tested sample matrices, indicating strong adaptability for complex biological backgrounds and multi-cancer methylation detection workflows. These results are based on internal validation studies.