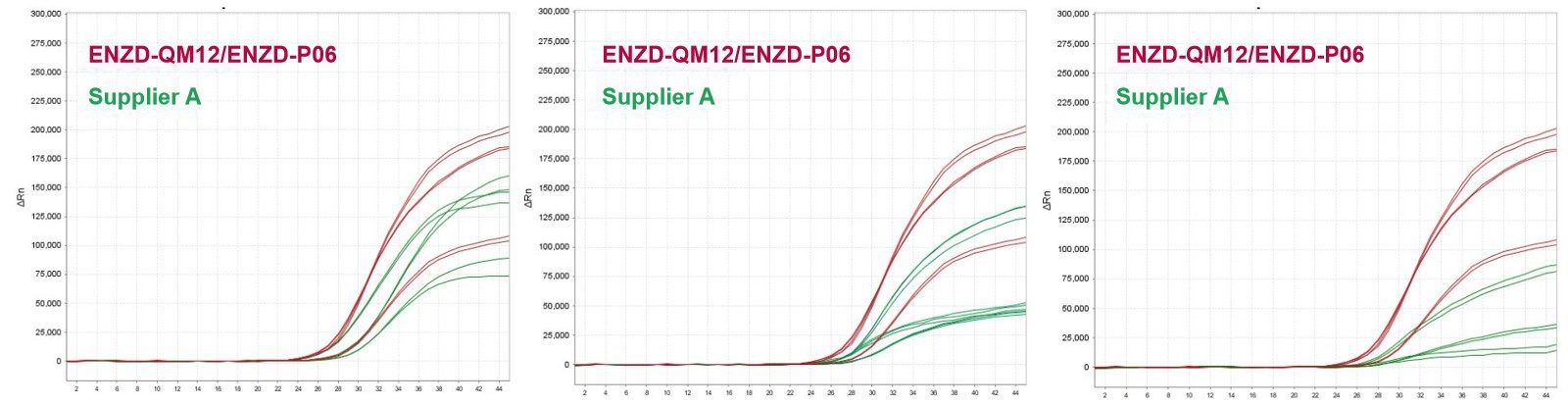
The performance of ENZD-QM12 was evaluated using bisulfite-converted DNA templates for methylation-specific qPCR analysis and compared with a commercial reference reagent under identical conditions.
Results showed that ENZD-QM12 delivered stronger fluorescence signals and higher amplification plateau levels, indicating improved reaction efficiency and more consistent amplification behavior in methylation detection workflows. This formulation is developed by Creative Enzymes and optimized for bisulfite-converted DNA analysis. Data sourced from internal validation studies evaluating amplification performance using bisulfite-converted DNA templates.
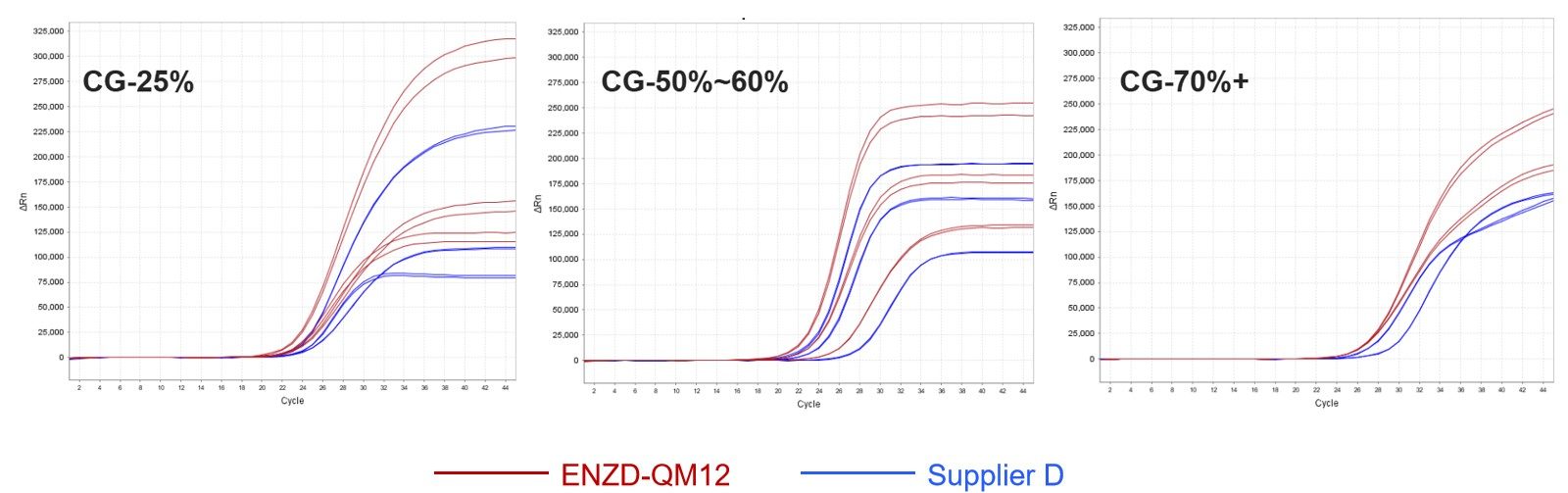
ENZD-QM12 was assessed using methylation-related CpG island targets with varying GC content ranging from low to high GC regions.
The system demonstrated stable amplification behavior across both low-GC and high-GC templates, indicating strong tolerance to GC variability and reduced amplification bias in methylation-specific qPCR applications. Data sourced from internal validation studies evaluating GC-content dependent amplification performance in bisulfite-converted DNA systems.
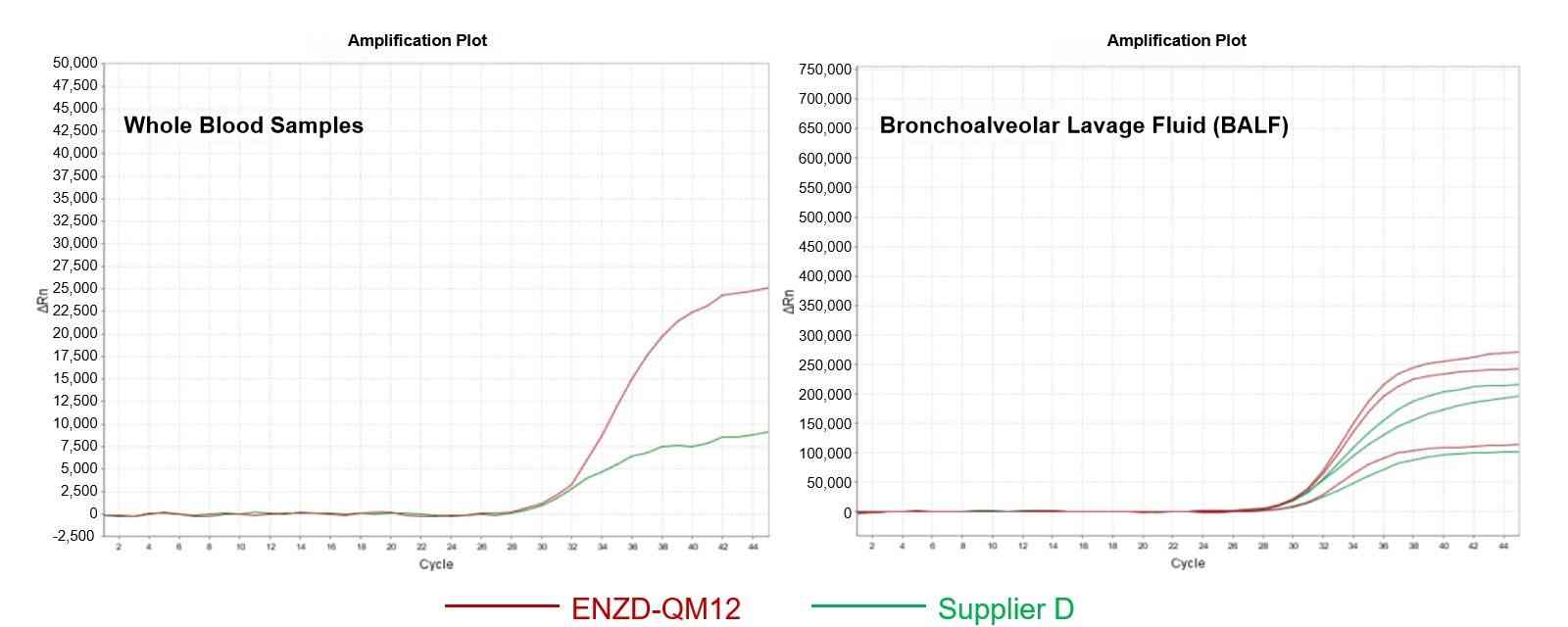
The applicability of ENZD-QM12 was evaluated using multiple clinically relevant bisulfite-converted sample types, including fecal extracts, exfoliated cells, blood-derived samples, and bronchoalveolar lavage fluid.
Across all tested matrices, ENZD-QM12 maintained stable amplification performance, indicating strong adaptability to complex biological backgrounds and supporting multi-cancer methylation detection workflows. Data sourced from internal validation studies evaluating sample matrix compatibility in bisulfite-converted DNA qPCR assays.